[ad_1]
Abstract
Introduction Researchers are increasingly developing algorithms that impact patient care, but algorithms must also be implemented in practice to improve quality and safety.
Objective We worked with clinical operations personnel at two US health systems to implement algorithms to proactively identify patients without timely follow-up of abnormal test results that warrant diagnostic evaluation for colorectal or lung cancer. We summarise the steps involved and lessons learned.
Methods Twelve sites were involved across two health systems. Implementation involved extensive software documentation, frequent communication with sites and local validation of results. Additionally, we used automated edits of existing code to adapt it to sites’ local contexts.
Results All sites successfully implemented the algorithms. Automated edits saved sites significant work in direct code modification. Documentation and communication of changes further aided sites in implementation.
Conclusion Patient safety algorithms developed in research projects were implemented at multiple sites to monitor for missed diagnostic opportunities. Automated algorithm translation procedures can produce more consistent results across sites.
Introduction
Health information technology shows promise for improving patient safety. Electronic health record (EHR) data are increasingly available and can prevent or detect potential patient safety events,1 thus providing knowledge to promote safety, learning and improvement. We previously developed electronic trigger (e-trigger) tools that query EHR databases to identify potential delays in follow-up of abnormal tests.2 Such algorithms can identify when a laboratory or radiology report suggests the need for additional testing, but appropriate follow-up has not occurred.3
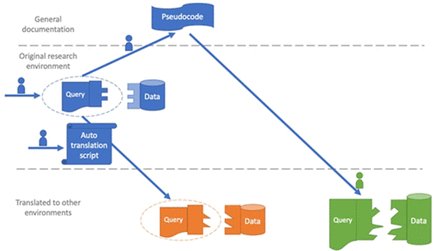
Patient safety algorithms developed through research must be implemented in clinical practice.4 However, there are no well-defined methods for implementation, and most studies do not make computer code available after publication,5 6 limiting opportunities to use algorithms clinically. Sharing code would improve replication, implementation and return on investment for research funding. A typical approach to reusing computer code in different institutions is to adapt each institution’s data to a common data model (CDM).7 Still, researchers invest much effort into algorithms that do not use CDMs. We believe that another alternative may advance the field: translate code and send it to sites with a supplemental description (figure 1).
Workflow of code translation from the research environment to multiple operational environments. In prior work, one team member developed structured query language algorithms to retrieve potential missed cancer follow-up cases from the Veterans Affairs (VA) research data warehouse (blue). For the present study, the team developed a script to translate the algorithms automatically to the VA operational data warehouse (orange), which is structured differently. The team also created pseudocode and documentation so that the algorithms could be translated to non-VA data warehouses (green), which have very different structures from VA.
We describe how researchers collaborated with multiple clinical sites to implement two algorithms that identify patients without timely follow-up of abnormal test results, warranting evaluation for lung or colorectal cancer. We also describe lessons learnt from the process.
Discussion
We successfully translated two patient safety algorithms from research to practice in multiple clinical sites, using a new approach: large-scale automated code translation rather than the typical method using a CDM.7 Lessons learnt include:
-
Write pseudocode with a complete value set listing for organisations with different data models.
-
Use source code control such as Git. Make code open to all sites.
-
Communicate frequently with sites receiving code.
-
Clinical personnel at each site should validate results.
Our approach is valuable when a research algorithm uses a non-standard data model; others can use the algorithm after translation to a new model. We expect centralised edits to decrease risk of errors and inconsistencies. Sending code to individual sites allows healthcare operations to benefit from our algorithms for missed tests concerning for cancer, by finding individual high-risk patients, notifying providers or measuring quality in a population. Apart from business reasons, there are scientific reasons for code sharing.9 A paper’s reviewers and readers should have access to the authors’ code to replicate the study, which they likely could not do from the methods section alone. Despite the push for research code sharing, a 2019 review showed 0 of 194 studies made analysis scripts available.5 Another showed that most studies decline to submit statistical code to a journal, or they include minimal documentation.6 Our online supplemental file 1 description is similar to the approach of Phenotype KnowledgeBase, a resource for sharing electronic phenotypes,10 but our code translation approach is unique, and our use of pseudocode for systems with different data structures is a strength.
Our work has several limitations. The adaptations required by our sites may not be desired by others. Second, the script that edits SQL code would have to be rewritten for other codebases. Nevertheless, the methodology of a programme automatically modifying another programme would be transferable and still save time. Third, since our focus was implementation, we did not quantify the benefit of pseudocode by assessing sites’ implementation before and after pseudocode, but this could be a topic for future research.
Conclusions
We describe a strategy to efficiently translate patient safety algorithms from research to practice in multiple health systems. We also provide generalisable lessons learnt. This approach impacts the care of individual patients, increases the return on investment of research funding, and potentially impacts long-term population health.
[ad_2]
Source link