[ad_1]
Discussion
This study has demonstrated the effectiveness of modern deep learning for clinical decision support. Evaluating transformer-based techniques against classical methods showed free-text in EHR contains untapped predictive information that can augment decision support tools. Evaluating on temporal splits captured repeat attendees, reflecting clinical reality,23 though similar performance was demonstrated when removing repeat patients.
Model performance analysis and comparison
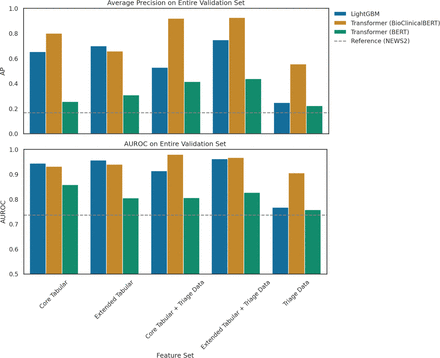
All machine learning models vastly outperformed NEWS, with models using free-text triage data outperforming those without (BioClinicalBERT tabular only AP, 0.80; with free text:,0.92). The best model, BioClinicalBERT with extended tabular+triage notes, outperformed NEWS across all performance metrics (eg, sensitivity 0.92 vs 0.13 at specificity 0.99). Pretraining on relevant in-domain data was crucial; standard BERT greatly underperformed their medical terminology-orientated counterpart, BioClinicalBERT (AP: 0.31 vs 0.66 on the same features). We surmise that the standard BERT pretraining corpus does not reflect the specialist language used in this task, limiting its performance.
BioClinicalBERT’s substantial performance gains with text features supports triage notes containing underused information, likely capturing patients’ social context and diagnostic severity, which is difficult to represent in structured fields. Previous work supports the notion that clinical acumen, as captured in free-text comments, can help predict patient outcome. The Nurse Intuition Patient Deterioration Scale has greater AUROC than NEWS, while in combination with NEWS, it can enhance rapid response systems.27 Likewise, the Dutch-Early-Nurse-Worry-Indicator-Score suggests that ‘worried’ nurses can identify deteriorating patients before their physiological vital parameters start to deteriorate.28 While our triage notes may not explicitly discuss prognosis or worry, there is clinical evidence justifying their inclusion.
The BioClinicalBERT model using only triage notes and demographics performed comparably to models built on tabular features and outperformed NEWS. This suggests it may be possible to embed a model at admission using the earliest available data, allowing early risk stratification before awaiting other clinical data.
Although BioClinicalBERT showed significantly improved discriminative ability over LightGBMs (see online supplemental table 4 for final LightGBM parameters), the simpler LightGBM models require less computation and are more interpretable, so it may be more viable in clinical settings when performing similarly well to transformers. With only tabular data tree-based models matched transformer performance, suggesting trees may be preferred when lacking text data.
Alongside improved discriminative ability, our proposed methods demonstrably reduced alert rate compared with NEWS; BioClinicalBERT with triage notes reduced mean daily alerts (figure 2). Adding tabular data further reduced the alert rate and increased AUROC and AP, indicating fewer unnecessary alerts.
Explainability and bias analysis
Using SHAP,29 we showed that complex models can be explained to clinicians, although with high computational cost. Figure 3 reveals that, without free-text triage notes, BioClinicalBERT relied more on primary admission diagnoses, presenting complaint and admission specialities, suggesting that this information is encapsulated within triage notes. Conversely, BioClinicalBERT incorporating triage notes placed greater importance on measured features (eg, vital signs); we hypothesise that this is because direct measurements cannot be inferred from triage notes. Interestingly, LightGBM models exhibited similar feature attributions regardless of free-text inclusion, suggesting limited free-text utilisation. In contrast to global explanations, we demonstrated how local explainability can provide patient-specific explanations to understand deterioration risk and guide patient management plan development.
Compared with NEWS, our models had lower I2 values across all sensitivity thresholds, indicating reduced bias. Generally, higher fidelity feature sets exhibited less bias than lower fidelity (figure 2e). However, this analysis is limited to our recorded protected characteristics. Future work should consider fine-grained data, such as socioeconomic and community context, which are known predictors of clinical risk,30 as language models can exhibit unfair bias.31
Implications for deployment in a clinical context
As acute care data collection is not standardised, we made as few assumptions about the data as possible.12
Together with the methods’ handling of missing data, this supports our models’ generalisability across EHR. We demonstrated that machine learning risk prediction can be easily applied across different feature sets, showing they can be deployed to different hospitals despite varying data collection standards/procedures. Without the rigid data requirements of existing techniques, our methods are easier to deploy across settings.
We intentionally avoided setting classification thresholds, instead measuring discriminative skill; setting thresholds carries clinical, operational and ethical considerations.25 All of our models can be tuned to balance false-positive and false-negative outcomes based on healthcare provider/regulator preference. We see the adoption of machine learning models in clinical practice as decision support tools rather than decision-making tools. However, this must be appropriately balanced to combat alert fatigue.30 Our analysis showed that this is possible, as all models achieved fewer average daily alerts (figure 2a) and higher clinical utility or net benefit (figure 2f) than NEWS at all but the highest sensitivities. If deployed to match NEWS sensitivity, we would raise fewer alerts while achieving the same level of care. For example, fixing BioClinicalBERT with all feature sets to a sensitivity of 0.32 (matching NEWS ≥5) achieves a positive predictive value of 0.85 versus 0.18 for NEWS. Alternatively, if the decision threshold is softened to match the alert rate of NEWS, BioClinicalBERT would identify cases that NEWS would miss.
Strengths and limitations
We believe this is the only large-scale evaluation of transformer-based models with free-text data as an EWS successor. We systematically examined how including free-text features improves model performance (increasing AP from 0.66 to 0.92), highlighting these untapped features’ usefulness. Importantly, we demonstrated that free-text notes alone contained sufficient predictive information to surpass existing EWS (AP ours, 0.92; AP NEWS, 0.12). Using explainability techniques, we demonstrated how explanations can elucidate important patient-level features, potentially increasing trust in the model and guiding clinical conversations.
Computing the generalised entropy index (I2), we compared the bias levels of our techniques against NEWS, showing our models yielded fairer distributions of benefit. However, data availability limited analysis to age, ethnicity and biological sex. Future research should consider other sources of bias such as socioeconomic status and free-text bias. Furthermore, our study contains data from a single site only. Data shift can affect the machine learning performance, and patient populations may vary significantly between hospitals;13 therefore, a multisite evaluation of our proposed techniques is warranted.
The use of free-text fields may differ between hospitals and requires further investigation; nomenclature, processes and data collection will differ between hospitals, possibly affecting model generalisability, necessitating a multicentre study. There were high rates of missing data, though reasons for this varied. Some were clinically meaningful, that is, the measurement was not clinically relevant. In other cases, values may not have been entered into the EHR correctly, perhaps because of operational pressures. We deliberately used models that can handle missing data, believing these yield techniques that are more applicable to real-world settings and allows for heterogeneity in features collected between hospitals. However, future studies should investigate the effect of missing data on the modelling process.
This study only showed the feasibility of using ML as an alternative to existing EWS. Prospective studies of our techniques are required to assess the impact of our models in clinical practice. These studies should consider factors such as usability and patient outcomes compared with existing EWS, together with patient and public involvement.
Comparisons with other studies
Previous machine learning models have been proposed as EWS replacements,32 but to our knowledge, ours is the first to include free-text data. Our LightGBM models using only tabular features achieved higher performance than similar studies16 (AP our model, 0.75; AP previous, 0.53), while our best transformer-based techniques vastly outperformed them (AP ours, 0.92,;AP previous, 0.53). Recent systematic literature reviews32 report that many studies fail to report suitable metrics for imbalanced classification (eg, F1 or F2 score), instead reporting the AUROC metric which we demonstrated is unsuited to imbalanced data. Direct comparisons with previous studies are difficult due to obscured discriminative power, different test sets and varying critical event definitions.32 Notably, few prior studies have compared directly to existing EWS.16 32 Unlike previous studies,32 we have demonstrated explainability techniques and evaluated bias.
[ad_2]
Source link