[ad_1]
Abstract
Background Despite the increasing availability of electronic healthcare record (EHR) data and wide availability of plug-and-play machine learning (ML) Application Programming Interfaces, the adoption of data-driven decision-making within routine hospital workflows thus far, has remained limited. Through the lens of deriving clusters of diagnoses by age, this study investigated the type of ML analysis that can be performed using EHR data and how results could be communicated to lay stakeholders.
Methods Observational EHR data from a tertiary paediatric hospital, containing 61 522 unique patients and 3315 unique ICD-10 diagnosis codes was used, after preprocessing. K-means clustering was applied to identify age distributions of patient diagnoses. The final model was selected using quantitative metrics and expert assessment of the clinical validity of the clusters. Additionally, uncertainty over preprocessing decisions was analysed.
Findings Four age clusters of diseases were identified, broadly aligning to ages between: 0 and 1; 1 and 5; 5 and 13; 13 and 18. Diagnoses, within the clusters, aligned to existing knowledge regarding the propensity of presentation at different ages, and sequential clusters presented known disease progressions. The results validated similar methodologies within the literature. The impact of uncertainty induced by preprocessing decisions was large at the individual diagnoses but not at a population level. Strategies for mitigating, or communicating, this uncertainty were successfully demonstrated.
Conclusion Unsupervised ML applied to EHR data identifies clinically relevant age distributions of diagnoses which can augment existing decision making. However, biases within healthcare datasets dramatically impact results if not appropriately mitigated or communicated.
Introduction
The increasing availability of electronic healthcare data has presented an array of opportunities to improve healthcare services by enabling data-driven decision-making.1–3 Despite this, and the discourse surrounding artificial intelligence (AI) and machine learning (ML), the adoption of data-driven tools within routine hospital workflows has remained limited. This study addressed two fundamental questions for using data-driven insights, derived from electronic healthcare record (EHR) data, using an example analysis. The first question concerns whether ML analysis is possible with EHR data and if so what kind. The second question concerns how the results of such ML analysis should be effectively communicated.
The use of EHR data to derive insight into healthcare processes is not new. Recently, a study used outpatient data along with k-means clustering to identify the most frequent diagnoses in a children’s hospital in China.4 However, numerous factors, including hospital processes and external socioeconomic influences can result in significant diversity across healthcare populations. As such, the extent to which any part of an ML pipeline or results are transferable remains uncertain. Furthermore, ML-based analysis has become increasingly accessible, with the introduction of easy-to-use APIs5 and a technology stack which has been abstracted away from companies and analysts.6 Additionally, the use of robust ’simple’ algorithms, for example, k-means clustering, has meant that producing reasonable findings based on ML analysis has been readily achievable. However, often the assumptions underpinning such analyses are not communicated and the results may therefore be over interpreted.
To address the first query, the problem of defining disease age clusters as a real-world use case, was considered. It is well-established that many diseases, while affecting a wide age range, have characteristic age distributions. Furthermore, several such distributions have already been described for specific conditions such as uveitis, asthma and thyroid disorders.7–9 Providing quantitative insights into these distributions would directly enable operational and policy level, data-driven decisions to be made, such as through accurate resource forecasting and planning. Furthermore, such analyses would enable clinicians to better understand the likelihood of presenting with different conditions at various ages and, for example, enable clinicians to understand the age atypicality of a given patient. Despite the clear advantages of understanding the age distributions and clusters across different diseases, the focus of the present study is predominantly on the modelling methodology within the context of healthcare, rather than any novelty of clinical insights. To address the second question, explicit focus was placed on the effect of various preprocessing decisions and how these could be effectively understood and communicated, for example, by examining the preprocessing of ICD-10 diagnosis codes. ICD-10 diagnosis codes assigned as part of a hospital’s billing process inform a large majority of statistical analysis and have been the subject of interest for recent research into applying ML to healthcare.10 Despite this, the inherent bias within the code assignments has seldom been addressed.
Discussion
The results presented have demonstrated the efficacy of using EHR data for deriving quantitative insights for clinical practice. This was demonstrated quantitatively, since a silhouette score17 of 0.15 was achieved and qualitatively, since the findings were broadly consistent with expected clinical and biological pathophysiologies. Given this, the result was considered as a validation of previous work using a completely independent dataset from a different population.4 Even accounting (partially) for the bias induced by the ICD-10 coding process, the learnt clusters were identified as grouping conditions predominantly affecting: young infants (ie, broadly between the ages 0 and 1); infants to young children (ie, broadly between the ages 1 and 5) with a peak in very early years; children (ie, broadly between the ages 5 and 13) and; teenagers and adolescents (ie, broadly between the ages 13 and 18).
However, care should be taken before generalising the findings of this study beyond the assumptions made in the analysis. ML methods generally assume the data on which the model was trained follows the same distribution as the data on which the model/insights from the model are being applied. While the general methodology of clustering diagnoses by defining age distributions may be applied elsewhere, applying the findings of this analysis to populations from different hospitals or populations, may result in invalid conclusions. Furthermore, the relationships presented here are associations and no suggestion is made regarding causality.
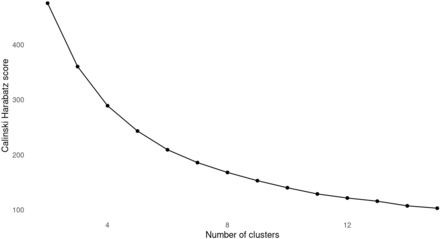
Future work could consider additional patient features as well as the use of other clustering methods. By their construction, ML methods assign different assumptions on the data which are required to hold for the clustering to be optimal and valid. For example, k-means clustering assumes the data forms independent spherical groups with constant radius.18 While the purpose of the current study was to demonstrate a proof of concept of the approach using routine EHR data and standard methods, given that the final clustering approach achieved a silhouette score of 0.15 (ie, less than 1), it may be possible to find superior separation of clusters with additional methods.
The preprocessing assumptions made when performing the modelling have been explicitly demonstrated to have a large effect on the results, an observation which has not generally been presented within the majority of applied ML for healthcare literature. Addressing uncertainty via altering the analysis hypothesis, while possible, required being extremely precise. Notably, the example of ‘Given a patient is aged X, what is the probability they will have condition Y?’ is not the only query a lay practitioner may have when presented with analysis defining patient age clusters. This suggested the need for further research into how ML tools and results can be better disseminated and reliably scaled and applied across healthcare organisations. Considering PedMap4 as an example, uncertainty resulting from a broad hypothesis could be reduced by tailoring the application views even further, to be focused on ‘clinical questions’ such as the one considered in this paper. It should be noted that the challenges associated with preprocessing ICD diagnosis codes are not unique to the analysis presented or dataset used and are prevalent throughout clinical informatic applications. Considering how to remove such biases and derive independent data representations that ‘truly’ reflect the holistic health status of patients (including diagnosis) is an interesting avenue of future work which may improve the results of the analysis presented in this paper.
Bayesian analysis has been generally conceived as difficult and subjective however, the results of the current study have demonstrated that even engaging with the Bayesian framework (without performing any Bayesian inferences) was intuitive and could prevent over interpretation of results. Rather than point estimates of clusters (such as those provided in PedMap), a distribution of clusters could be provided as output, such as that provided in figure 2. In the absence of techniques to directly remove the bias arising from preprocessing assumptions, a Bayesian inference approach to defining this uncertainty would be an interesting next step. Additional interesting avenues of future work might be to focus on expanding the ‘plug and play’ ML APIs to allow practitioners to incorporate Bayesian uncertainties over preprocessing steps, with only an introductory understanding of Bayesian inference.
[ad_2]
Source link