[ad_1]
Abstract
Background Chronic kidney disease (CKD) is a global health concern characterised by irreversible renal damage that is often assessed using invasive renal biopsy. Accurate evaluation of interstitial fibrosis and tubular atrophy (IFTA) is crucial for CKD management. This study aimed to leverage machine learning (ML) models to predict IFTA using a combination of ultrasonography (US) images and patient biomarkers.
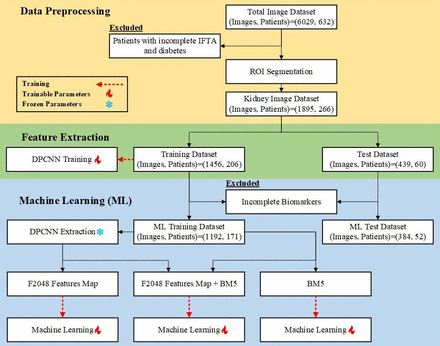
Methods We retrospectively collected US images and biomarkers from 632 patients with CKD across three hospitals. The data were subjected to pre-processing, exclusion of sub-optimal images, and feature extraction using a dual-path convolutional neural network. Various ML models, including XGBoost, random forest and logistic regression, were trained and validated using fivefold cross-validation.
Results The dataset was divided into training and test datasets. For image-level IFTA classification, the best performance was achieved by combining US image features and patient biomarkers, with logistic regression yielding an area under the receiver operating characteristic curve (AUROC) of 99%. At the patient level, logistic regression combining US image features and biomarkers provided an AUROC of 96%. Models trained solely on US image features or biomarkers also exhibited high performance, with AUROC exceeding 80%.
Conclusion Our artificial intelligence-based approach to IFTA classification demonstrated high accuracy and AUROC across various ML models. By leveraging patient biomarkers alone, this method offers a non-invasive and robust tool for early CKD assessment, demonstrating that biomarkers alone may suffice for accurate predictions without the added complexity of image-derived features.
Introduction
Chronic kidney disease (CKD) causes significant morbidity and mortality worldwide, with global prevalence rates of 9.1% and 697.5 million cases.1 It is characterised by irreversible damage to the renal tissue, which can ultimately lead to end-stage kidney disease, resulting in a substantial economic burden.2 A previous study showed that accurate assessment of renal interstitial fibrosis and tubular atrophy (IFTA) is crucial for diagnosing and managing CKD.3 IFTA severity is conventionally assessed through renal biopsy, which remains the gold standard for obtaining detailed histopathological information. This procedure provides direct visualisation and quantification of IFTA but is time-consuming and subject to inter-observer variability. Additionally, renal biopsy is an invasive procedure, making it unsuitable for all patients.4
Medical ultrasonography (US) is a crucial diagnostic tool for kidney or ureteral structural diseases, which measures imaging parameters that provide vital information regarding renal function. Previous studies have explored different parameters such as kidney size, cortical thickness and cortical echogenicity to estimate changes in estimated glomerular filtration rate (eGFR).5–12 A previous study revealed a significant positive correlation between eGFR and the mean renal length (r=0.66) and mean cortical thickness (r=0.85).5 Additionally, changes in the texture of the renal tissue on US images can also suggest changes in renal function. Nevertheless, interpreting US images of the kidney requires extensive training for clinicians, and the results may lack objectivity due to subjectivity.
Recently, artificial intelligence (AI) has emerged as a promising tool for predicting pathological results by leveraging data acquired through non-invasive methods.13 14 Deep learning using convolutional neural networks (CNNs) has also demonstrated good performance in the analysis of medical US images. In a previous study, CNNs were used to grade the severity of inflammation in the long head of the bicep tendon.15 Moreover, one study relied on CNNs and consecutive comprehensive non-stress echocardiography to predict cardiac function in patients to better understand ageing and prevent cardiovascular diseases.16 Similarly, a deep-learning algorithm using kidney US images of a single centre accurately quantified IFTA with 90% accuracy, indicating its potential as a non-invasive first-line investigation for kidney-disease assessment.17 However, there is a lack of studies conducted across multiple medical centres in leveraging AI for predicting pathological results through non-invasive data acquisition methods.
Our research aimed to leverage AI to predict the stage of IFTA using a combination of demographic data, laboratory results and renal US images across diverse medical centres. Specifically, we focused on five key clinical biomarkers: age, sex, eGFR, serum albumin and kidney size. Renal US provides a non-invasive and readily available imaging modality that can capture detailed structural information regarding the kidneys. We compared three approaches: using only biomarkers, relying solely on US images and integrating both modalities for a comprehensive analysis.
Discussion
The development and validation of our AI algorithm involved a multistep process to ensure robustness and accuracy for both image-level and patient-level classifications. The use of biomarkers alone yielded optimal results in our analyses, demonstrating that the structural insights from imaging did not significantly enhance predictive accuracy. The use of fivefold cross-validation helped minimise bias and optimise model performance, leading to high accuracy and AUROC scores across various classifiers.
Previous studies investigated the correlation between renal US images and IFTA scores.29 These studies found that sonographic parameters such as kidney length, echogenicity and parenchymal thickness demonstrated only weak-to-moderate relationships with interstitial fibrosis or tubular atrophy, with the highest Spearman correlation coefficient reaching 0.35. In another study, researchers proposed an AI model to predict IFTA scores based on renal US images.17 The UNet architecture was employed for US image segmentation, and the VGG19 and XGBoost models were used for feature extraction and image classification, respectively. The prediction model achieved a performance ranging from 0.8037 to 0.8927 in terms of accuracy, precision, recall and F1 score. However, this study relied on retrospective data from a single centre.
One of the major strengths of our study is the comprehensive dataset, which included a large number of images from a diverse patient cohort across multiple hospitals, enhancing the generalisability of our model. Additionally, our approach combines the analysis of US images with patient biomarkers to provide a valuable layer of information that improves the predictive power of our models. This method demonstrated a robust performance across various ML techniques, consistently achieving an AUROC >90%. While it cannot entirely substitute human interpretation and clinical expertise, it can serve as a valuable tool for assisting less-experienced physicians in providing accurate interpretations. Future research should focus on investigating the long-term impact and cost-effectiveness of this technology, as well as exploring effective ways to integrate it into existing clinical practices.
Despite these promising results, our study has several limitations. First, the retrospective nature of the study may have introduced biases related to data collection and patient selection, which may have affected the generalisability of our findings. Second, the exclusion of patients with diabetic nephropathy, while necessary to avoid misleading the model, may limit the applicability of our results to a broader population, particularly those with comorbid conditions. Third, the manual review process for image quality is subject to human error and variability, which can affect the consistency of the data used for training and validation. Fourth, while our model demonstrated high performance, with AUROC >90% in the independent test set, it needs to be validated in external cohorts to confirm its robustness and generalisability beyond the studied population. Fifth, the lack of standardised pathology guidelines across hospitals in our country posed a significant limitation. As each pathology department follows different guidelines, detailed parameters such as IFTA foci density could not be consistently obtained or included in this study. Finally, follow-up data were not available in our study to assess the model’s capacity to predict long-term outcomes such as end-stage kidney disease (ESKD). Future studies should include follow-up information to evaluate the model’s potential for predicting ESKD and its broader clinical utility.
Conclusions
Our study presents a robust AI-based approach for IFTA classification that relies on patient biomarkers alone, demonstrating that image features do not significantly enhance predictive performance. The model demonstrated high performance across various ML techniques, consistently achieving AUROC >90%. This method, based on biomarkers alone and developed using a comprehensive dataset from multiple hospitals, has the potential to enhance early determination of IFTA without biopsy, avoiding unnecessary complexity from image features. Future studies should focus on validating the performance of the model in external cohorts to ensure its generalisability and applicability to diverse clinical environments.
[ad_2]
Source link