[ad_1]
Discussion
Even though the crucial role of data in clinical practice, healthcare research and quality assessment cannot be questioned, data collection from medical records is a time-consuming and error-prone task that may shift physician time allocation away from direct patient care.1–3 Automating these tasks with the use of LLMs can help unburden physicians from data collection tasks and increase the quantity and quality of time spent with patients. LLMs can be used in data collection from unstructured and semi-structured medical records, such as referral letters, discharge summaries, radiology reports, pathology reports, and blood tests, and, thus, can assist physicians and other healthcare professionals directly with the systematic processing of previous medical information, and indirectly with clinical decision-making. Furthermore, such implementation of LLMs can help leverage medical data in under-resourced hospitals, and assist or undertake data collection for newly established and previously non-existent databases, processing large quantities of unstructured and semi-structured medical notes stored in electronic health records.12
The process of manually extracting data from medical records is rather tedious than complicated and sophisticated, performed mainly by junior doctors or other adequately trained healthcare professionals. Modern LLMs have exhibited a remarkably high, human-level performance on various professional and academic benchmarks in numerous domains, including medicine.8 In an analysis by Nori et al, GPT 4 exceeded the passing score of USMLE by over 20 points in a zero-shot setting.13 In view of this notable performance of GPT 4 in medical licensing examinations, we hypothesised that GPT 4 and other modern LLMs might be able to reliably perform the relatively low-complexity task of entity extraction from and binary classification of text from medical notes.
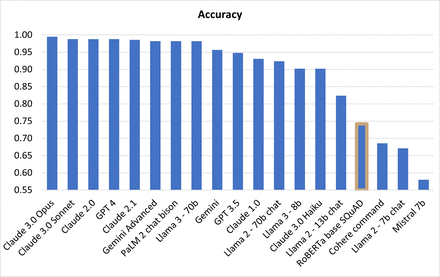
Eight LLMs showed an accuracy of over 0.98. Claude 3.0 Opus, exhibited the highest performance overall, returning a correct value for all but two of the total 450 requested values and having the highest performance metrics of all LLMs assessed in this study, with an accuracy over 0.99. The second best performance in terms of number of correct values and accuracy, exhibited Claude 3.0 Sonnet, Claude 2.0 and GPT 4, all of which returned five false values each. Following, were Claude 2.1, Gemini Advanced, PaLM 2 chat bison and Llama 3-70b, each of which returned six, eight, eight, and eight false values, respectively. All false values returned by Claude 3.0 Opus, Claude 2.0, GPT 4, Claude 2.1, PaLM 2 chat bison, and Llama 3-70b were binary classifications, whereas Claude 3.0 Sonnet and Gemini Advanced additionally returned two and one false value, respectively in entity extraction. None of the eight highest performing LLMs returned non-requested/unmatched values. Except for two missing values for PaLM 2 chat bison, and two misclassifications to values other than the ones explicitly requested in the LLM prompt for Gemini Advanced, no missing values and misclassifications were returned by Claude 3.0 Opus, Claude 2.0, Claude 3.0 Sonnet, GPT 4, Claude 2.1, and Llama 3-70b.
The abovementioned eight highest performing LLMs exhibited higher performance than the baseline RoBERTa model with at least 32% higher overall accuracy, at least 140% higher recall and at least 70% higher F1 score than the baseline model. Regarding precision, Claude 3.0 Opus, GPT 4 and Gemini Advanced exhibited a marginally higher (maximally 0.4%), and Claude 3.0 Sonnet, Claude 2.0, Claude 2.1, PaLM 2 chat bison and Llama 3-70b a slightly lower performance (maximally −4.1%) than the baseline RoBERTa model. Finally, regarding multiple-run response consistency, Claude 2.0 exhibited the highest performance, with a perfect response agreement over all three runs, whereas six of the eight highest performing LLMs showed a marginally higher (maximally 1.2%), and only Gemini Advanced a marginally lower (−0.3%) consistency than the baseline RoBERTa model.
Different LLMs have different context lengths, and recent LLMs have increasingly larger context lengths, however, concerns exist about LLM performance in longer context texts. In a study analysing LLM performance on multi-document question answering and key-value retrieval, it was found that performance can degrade significantly when changing the position of relevant information in the input text. Particularly, the highest performance was often observed when information was placed at the beginning or end of the input text, and the lowest performance when information was placed in the middle of long texts, even for explicitly long-context LLMs.14 Another study, assessing long-context LLM performance on different input text lengths, found large performance drops as the input text length increases, indicating that the effective context length can be lower than the claimed context length, with some LLMs exhibiting larger performance drops than others.15 As a result, increasing clinical note length may have a negative impact on the performance of several LLMs.
The accuracy of a diagnostic model varies directly with disease prevalence, and the upper and lower bounds of accuracy are determined by the model’s sensitivity and specificity. In a population with a disease prevalence of 100%, the accuracy of a model equals its sensitivity; in a population with a disease prevalence of 0%, accuracy equals specificity. Between the bounds determined by sensitivity and specificity, accuracy varies directly (linearly) with disease prevalence. If disease prevalence equals 50%, a model’s accuracy is exactly midway between its sensitivity and specificity.16 Consequently, for rare conditions with low prevalence LLM accuracy is expected to approximate LLM specificity. We calculated LLM specificity and LLM classification accuracy, and as expected, due to the 50% prevalence of each examined complication, the LLM classification accuracy was the exact mean of the LLM recall and LLM specificity (online supplemental material).
An analysis of the accuracy of manual data collection from electronic health records observed an average transcription error rate of 9.1% per patient dataset for retrieving a dataset containing 27 variables.17 Even higher extraction error rates, up to 50% in some cases, were found in a systematic review assessing the frequency of manual data extraction errors in the setting of meta-analyses.18 In our study, none of the evaluated LLMs exhibited a completely correct performance; however, considering the high transcription error rates observed in some studies, the performance of the highest performing LLMs, should be viewed as outstanding.
LLMs have shown better performance in few-shot as compared with zero-shot evaluations.13 However, in this analysis, only a zero-shot evaluation was performed, as this better approximates how human evaluators process medical notes in everyday practice. Another issue occasionally occurring when using LLMs is ‘hallucinations’ or confabulated responses, where the model’s response does not seem to be justified by text input and training data.19 In our study, some LLMs exhibited a few confabulated responses, where non-requested or values unmatched to the requested entity extraction and classification tasks were observed in the LLM output.
A major issue complicating the use of LLMs for data extraction from real patient records in a real-world setting, is the submission of sensitive patient data to out-of-hospital computer infrastructure, which due to the associated patient data security risks, requires approval by the hospital data governance board and local ethics committee, and presents a serious, if not insurmountable, challenge. Even though this issue concerns LLMs running in out-of-hospital cloud infrastructure, local installations of open-source LLMs might be able to deal successfully with this issue, although running an LLM locally can be computationally intensive and require significant computational resources. Deidentifying and date-shifting clinical data, as used by Sushil et al in a study of LLM-based classification of breast cancer pathology reports, can be another solution to this issue, even though deidentifying discharge summaries and referral letters could be technically challenging.10
Even though some LLMs exhibited an impressively high performance in data extraction from medical notes, the results of this study should still be interpreted with caution. The present analysis has been based on the input of relatively short-sized synthetic medical notes, on a small number of entity extraction and binary classification tasks, with no assessment of classification performance for categorical variables with more than two-level responses. Furthermore, as the clinical notes were drafted from only one domain expert, text variability can be expected to be relatively limited, which might further affect the generalisability of these results to different institutions, regions and countries with different styles and abbreviations in documentation. Although noise consisting of typos, and abbreviations requiring context-dependent disambiguation was added to the synthetic text to better simulate healthcare practice, the clinical notes might still be over-processed and not fully representative of real-world text. Finally, this study assessed LLM performance only on data extraction from medical notes related to the postoperative course after cardiac surgery, and for a small number of related variables/complications, therefore, the generalisability of these results to other medical specialties or other medical contexts might be limited, and warrants further research for LLM performance assessment in other use cases. The key point regarding the use of LLMs for data extraction from medical records, is that as is the case with humans, LLMs are limited to identifying only what is documented in these medical records.
However, the strengths of this study should be also emphasised. Even though non-exhaustive in the number of LLMs included, our study assessed a large number of LLMs, evaluating the performance of 18 LLMs and comparing their performance against a fine-tuned question answering version of RoBERTa base, a nowadays less sophisticated transformer-based model; to our knowledge, our LLM list comprises most known closed- and open-source LLMs. Moreover, a model response consistency assessment was performed over three iterations of the same prompt for each model. Although the text entered in the LLM prompts was synthetic, it was carefully drafted and evaluated by domain experts to represent medical notes commonly found in clinical practice; in addition, it was class-balanced in order to assess better the discriminatory ability of each LLM for both positive and negative classes, desired attributes rarely present in real-world medical records; finally, to increase the robustness of the analysis, noise was introduced in the text. As a consequence, the results of this study are suggestive of the performance of the evaluated LLMs in a real-world setting.
[ad_2]
Source link