[ad_1]
Discussion
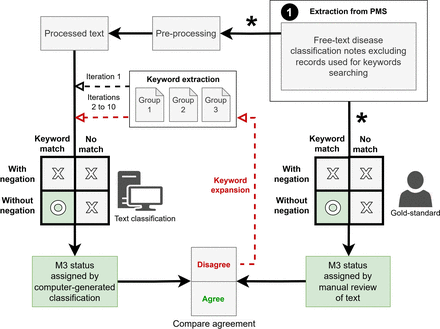
This study highlights the importance of incorporating unstructured free-text data from general practice EHRs to provide a more accurate representation of the burden of long-term conditions. Our analysis, which represents the first large multipractice NZ study of MM in general practice, revealed a substantial number of multimorbid patients whose conditions were not captured in Read coded data. By augmenting structured data with free-text analysis, our ‘R_M3Text_Classification’ procedure improved the identification of M3 index conditions, achieving very high specificity, sensitivity, positive predictive value and F1-score.
The sensitivity of the first iteration was 72.6%. This initial value was primarily due to overdetection of conditions that were acute or short term, related to family history, or resulted from routine screening. For example, people with gestational diabetes, a family history of gynaecological cancer, negative HIV or breast cancer screening tests and previous benign breast biopsy results were often misclassified as having a long-term condition. To improve sensitivity while minimising false positives, the programme was refined to accurately identify when keywords indicated conditions were not long term. The highly variable nature of free-text, particularly in a database sourced from multiple clinicians, meant that it was not possible to identify all permutations of phrasing. Despite these challenges, our methodology achieved consistently high specificity—minimising false positives—which is crucial for accurate condition identification in general practice.
Two clinical experts collaboratively developed key terms, resolved ambiguities and refined the classification rules, while our statistical expert translated those decisions into logical programmable steps. This iterative process has demonstrated that even a relatively straightforward text classification approach can yield robust performance without the resource-intensive requirements of advanced natural language processing (NLP) techniques.
Our findings align with similar studies that aimed to harness unstructured free-text data from EHRs to enhance disease classification and management. Recent research15–18 employing advanced NLP and machine learning techniques has successfully analysed unstructured patient medical records, enabling effective identification of patients with specific diseases. However, these methods often necessitate extensive computational resources and large datasets for training, which can limit their applicability in resource-constrained settings. In contrast, our study demonstrated that a more straightforward text classification approach, such as the ‘R_M3Text_Classification’ procedure, can yield high performance metrics without the complexities of NLP techniques, particularly in general practice free-text data. In line with recent work by Hossain et al,19 which highlighted challenges with misclassification due to ambiguous phrasing and negation in free-text entries, our study reinforces the notion that while sophisticated algorithms can enhance accuracy, simpler methods can also be effective when tailored to the specific characteristics of the data. These findings suggest that our approach may serve as a practical alternative for general practice settings, where clinician time and resources are often limited.
While our study yielded promising results, it is important to acknowledge limitations such as dependence on the quality of free-text records and lack of specific keywords for some conditions, particularly when the condition descriptions are broad. For example, it was difficult to identify keywords for venous insufficiency and uncomplicated hypertension that were specific enough to avoid false positives. Another limitation arises from the presence of pseudo-negations, such as double negatives (eg, not ruled out) or ambiguous negations, which can pose challenges for the procedure and potentially lead to misclassification.
While challenges remain, including dependency on the quality of free-text records and the difficulty of capturing all phrasing variations, our study provides a practical alternative for general practice settings. For example, consider a patient whose coded records indicate atrial fibrillation and coronary disease. However, on incorporating free-text entries from the same records, we identified that the patient had also developed heart failure—a condition not captured by structured data alone. This case exemplifies the broader implications of our approach, demonstrating that augmenting structured data with free-text analysis enhances MM detection and provides a more accurate representation of patients’ health status. Studies such as Owen et al20 have shown that the temporal sequence of MM diagnoses can significantly affect life expectancy, underscoring the clinical importance of capturing the full disease burden for optimal decision-making. Integration of our procedure into the PMS could improve medical documentation precision, enhance clinical decision-making and ultimately support better patient outcomes and health policy decisions. External validation across diverse patient populations is needed to further assess the applicability and effectiveness of this approach. This would provide valuable insights into its functionality in different general practice settings.
[ad_2]
Source link